Generalized Pythagoras Trees
for Visualizing Hierarchies

for Visualizing Hierarchies
Fabian Beck, Michael Burch, Tanja Munz, Lorenzo Di Silvestro, and Daniel Weiskopf. Generalized Pythagoras Trees for Visualizing Hierarchies. In IVAPP '14 Proceedings of the 5th International Conference on Information Visualization Theory and Applications. 2014 (to appear).
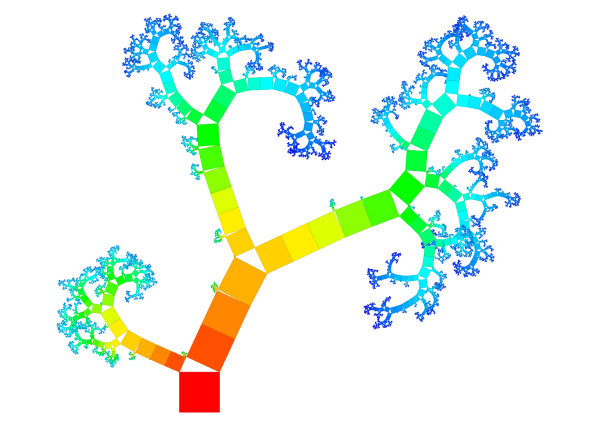
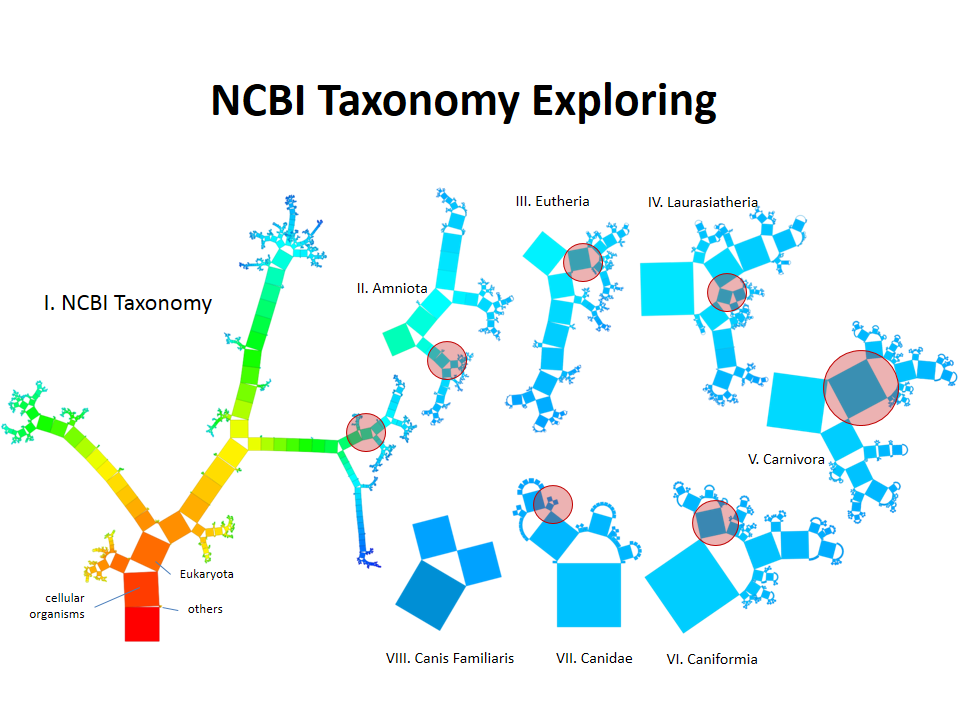
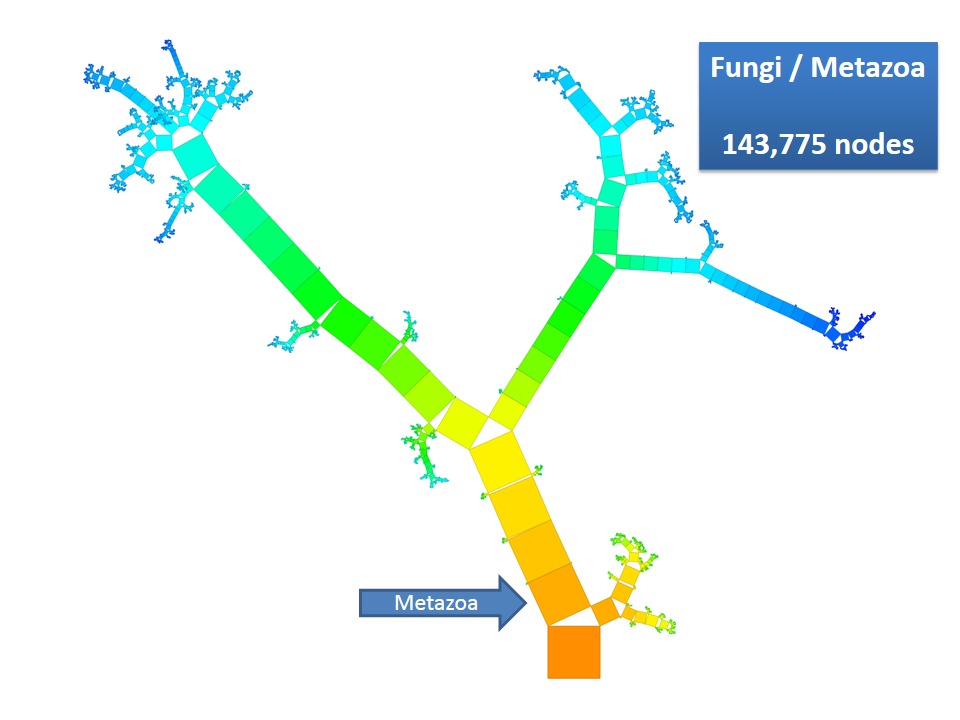
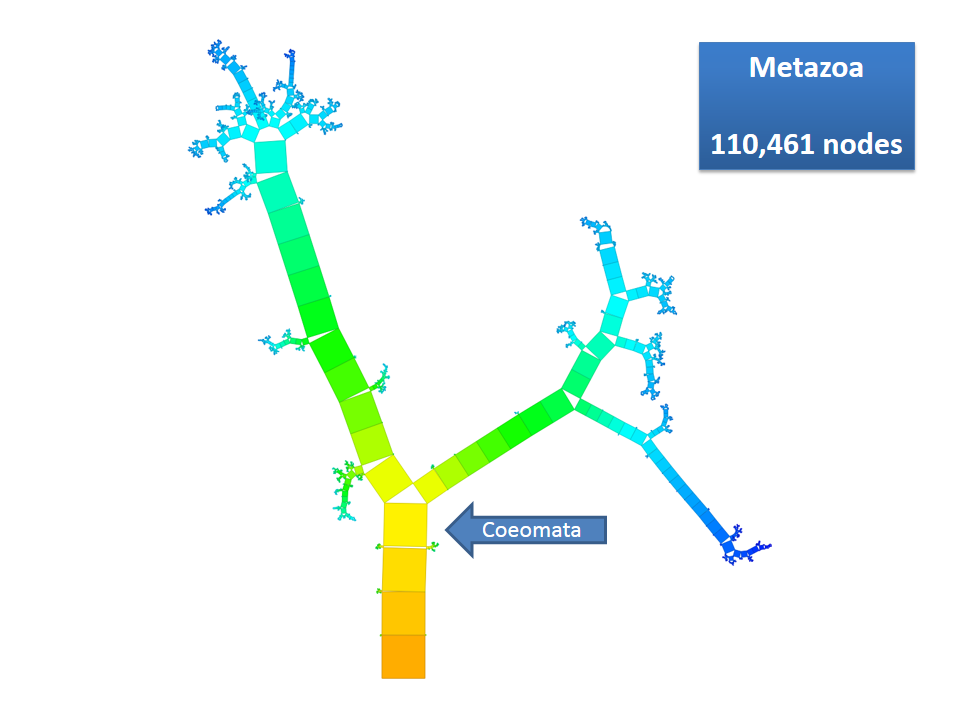
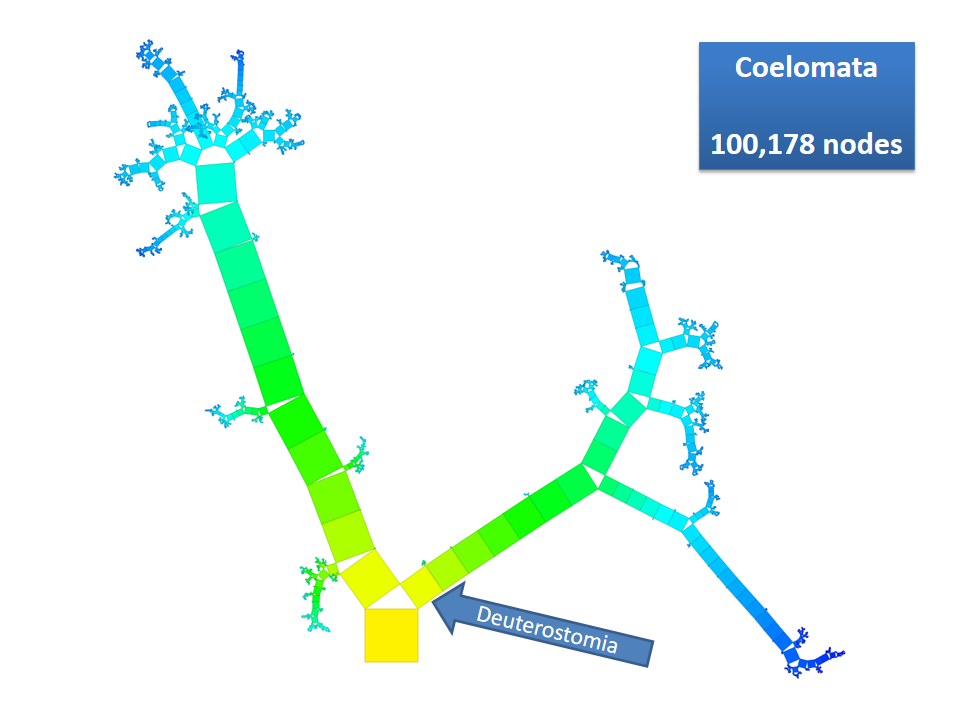
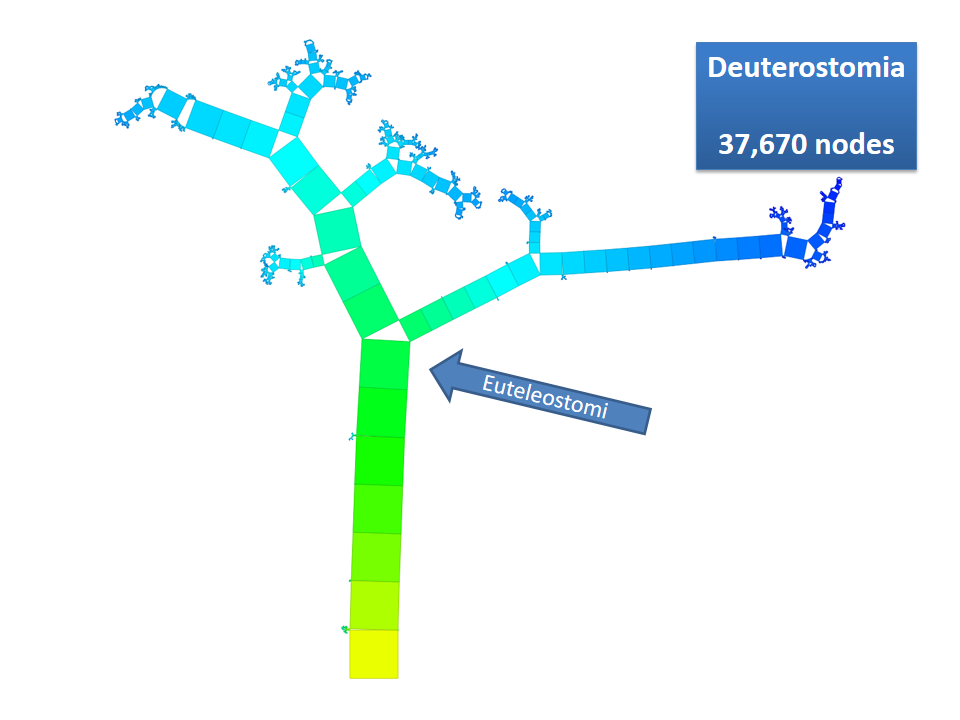
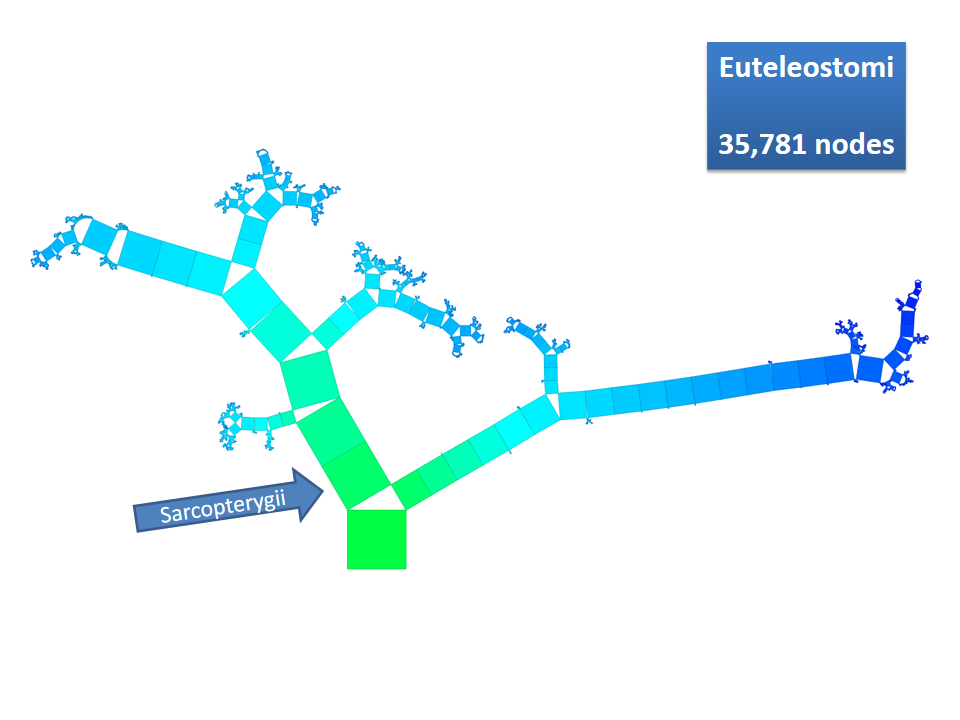
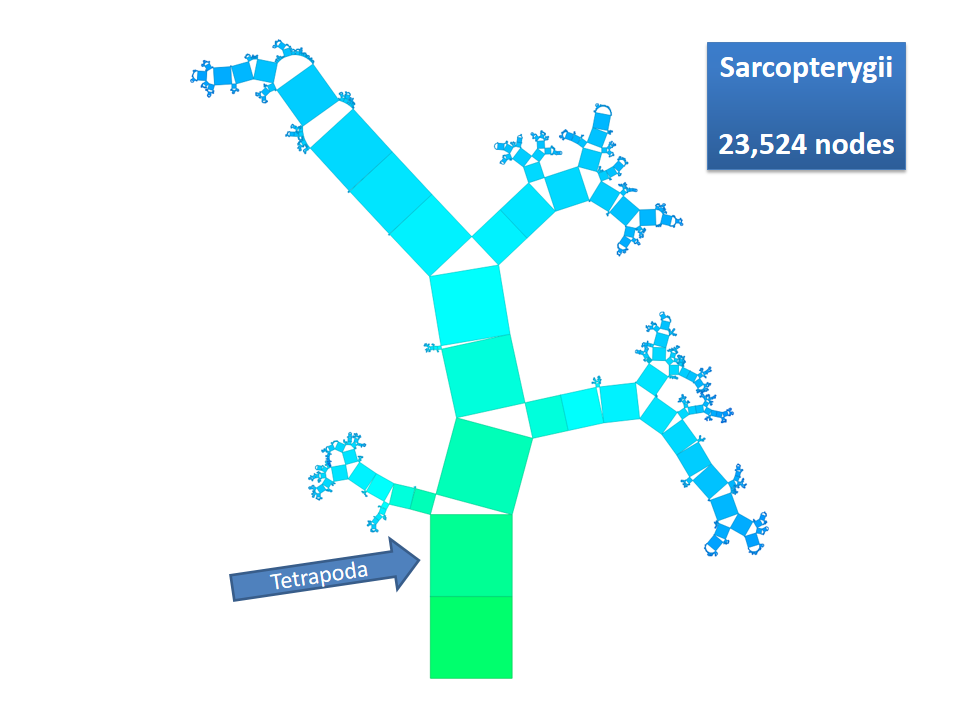
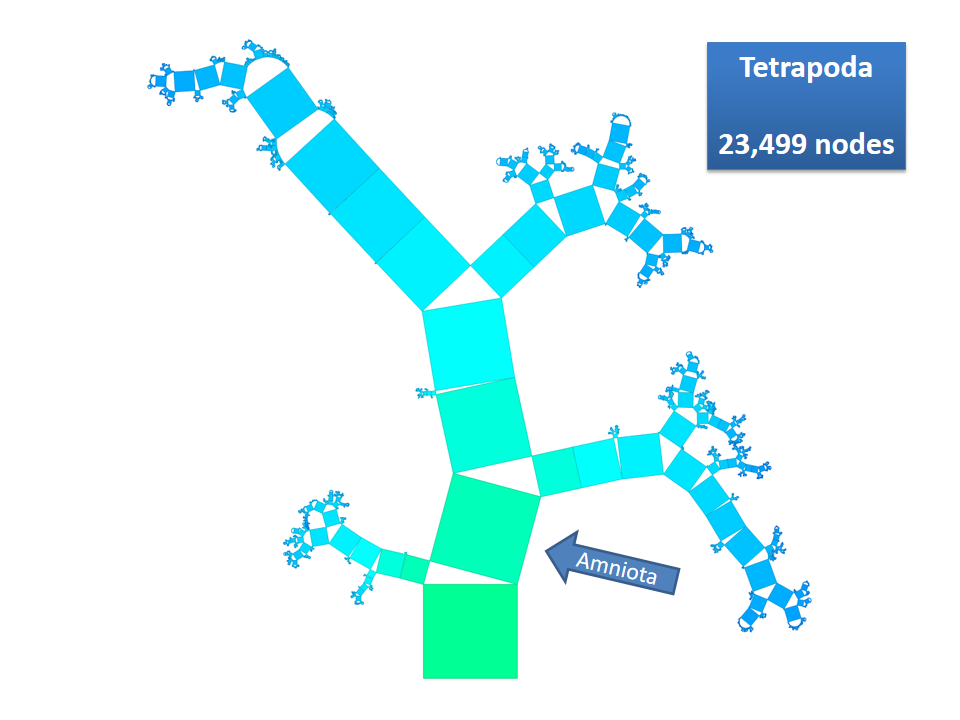
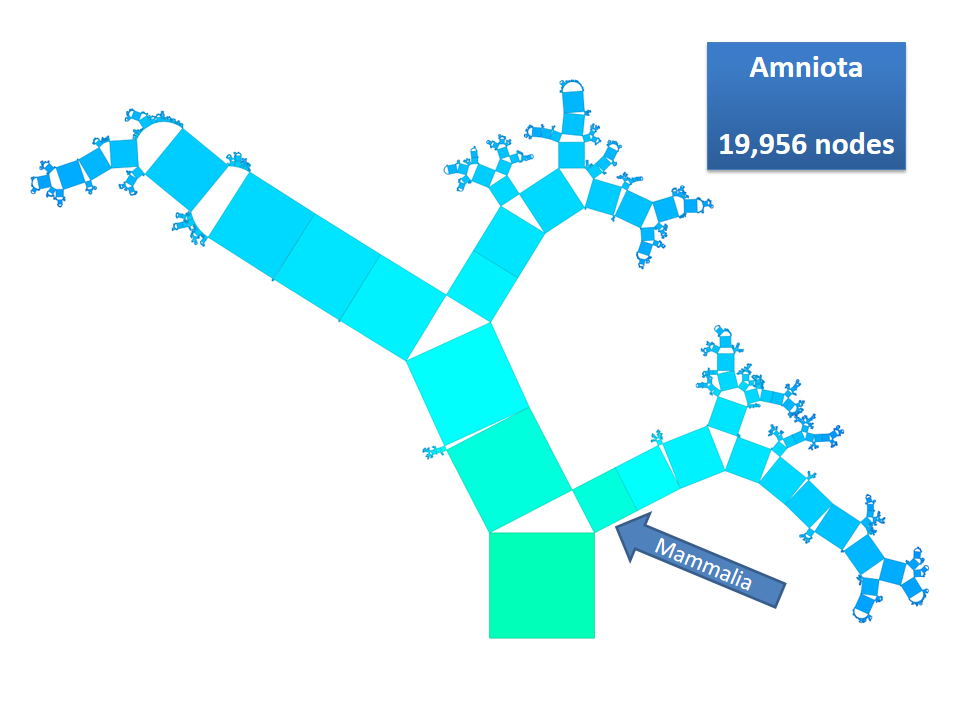
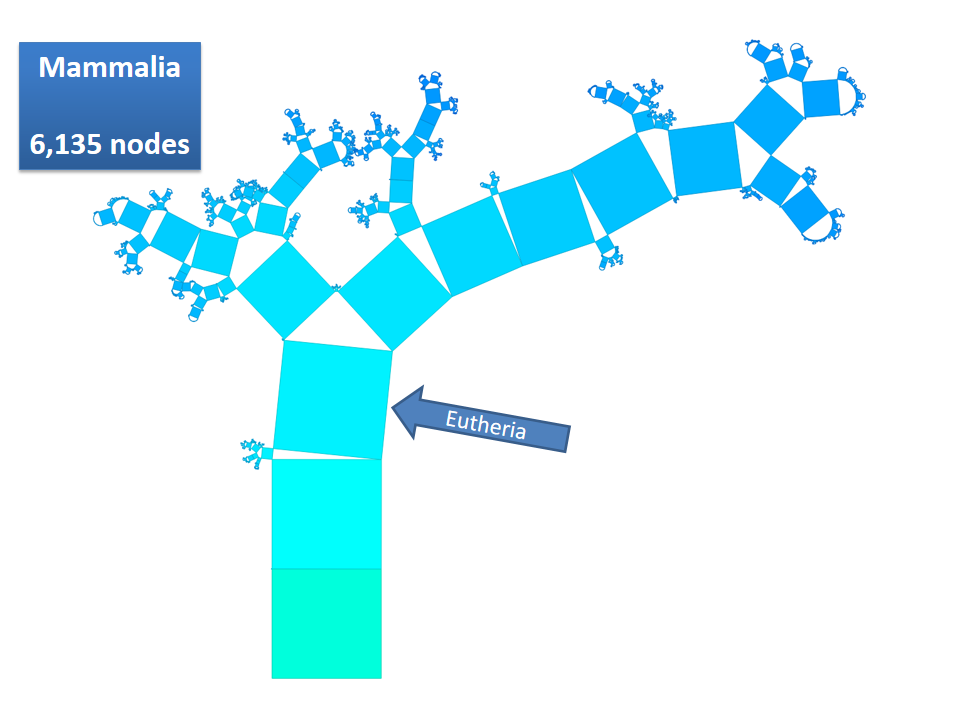
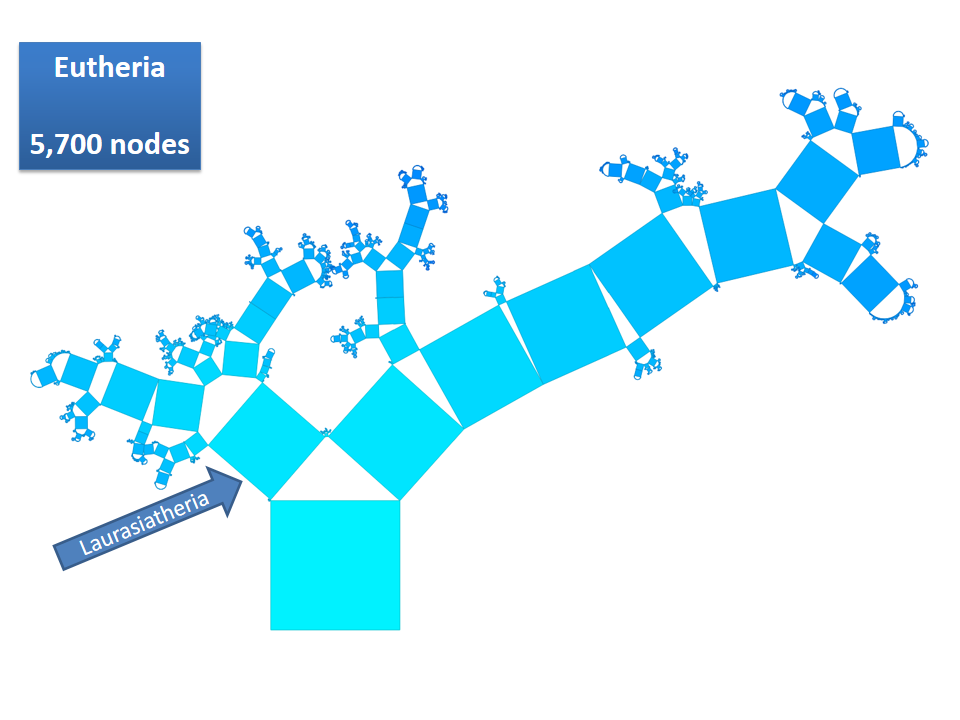
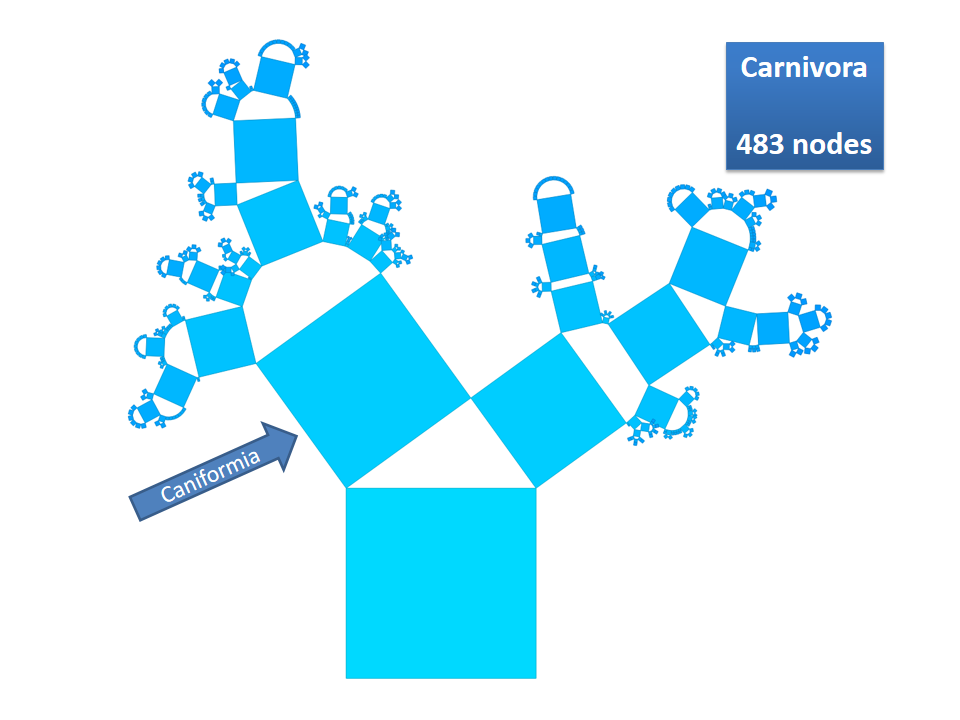
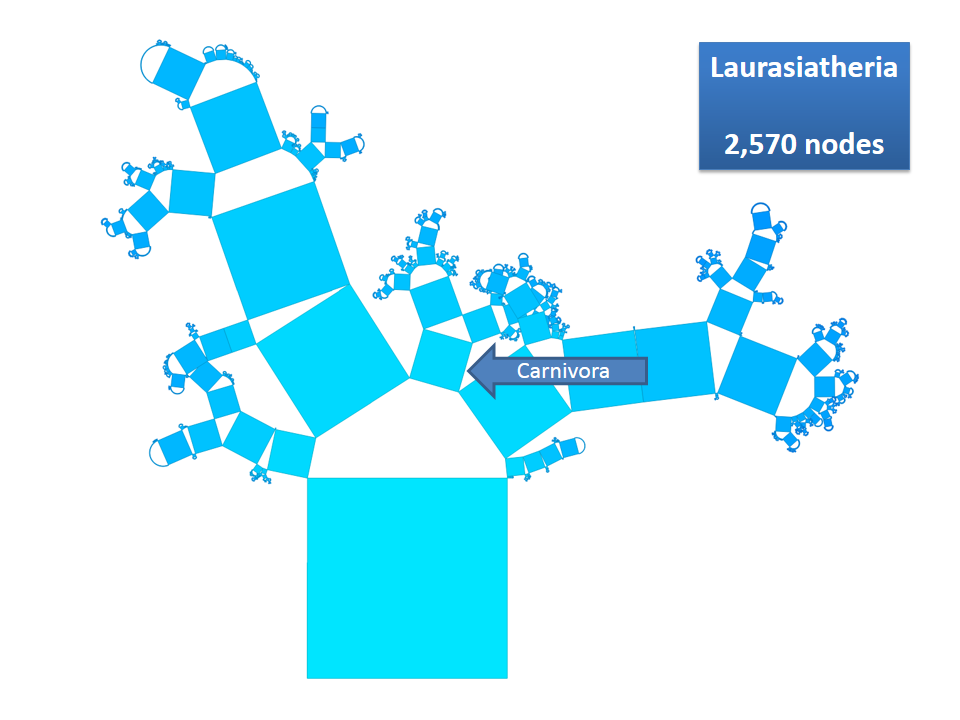
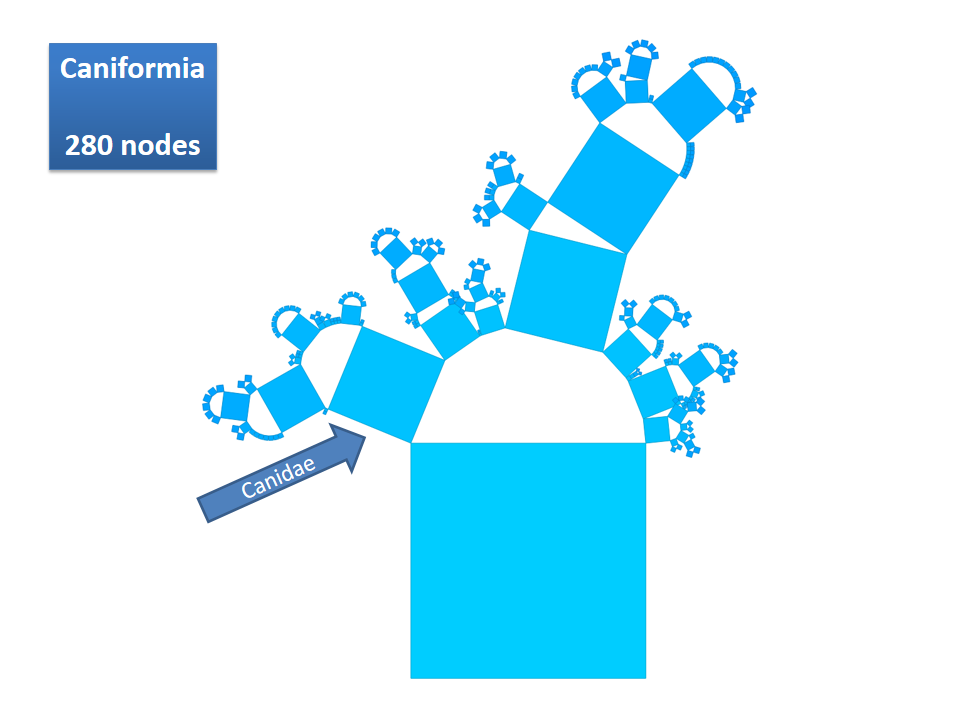
Pythagoras Trees are fractals that can be used to depict binary hierarchies. But this binary encoding is an obstacle for visualizing hierarchical data such as file systems or phylogenetic trees, which branch into n subhierarchies. Although any hierarchy can be modeled as a binary one by subsequently dividing n-ary branches into a sequence of n-1 binary branches, we follow a different strategy. In our approach extending Pythagoras Trees to arbitrarily branching trees, we only need a single visual element for an n-ary branch instead of spreading the binary branches along a strand. Each vertex in the hierarchy is visualized as a rectangle sized according to a metric. We analyze several visual parameters such as length, width, order, and color of the nodes against the use of different metrics. The usefulness of our technique is illustrated by two case studies visualizing directory structures and a large phylogenetic tree. We compare our approach with existing tree diagrams and discuss questions of geometry, perception, readability, and aesthetics.
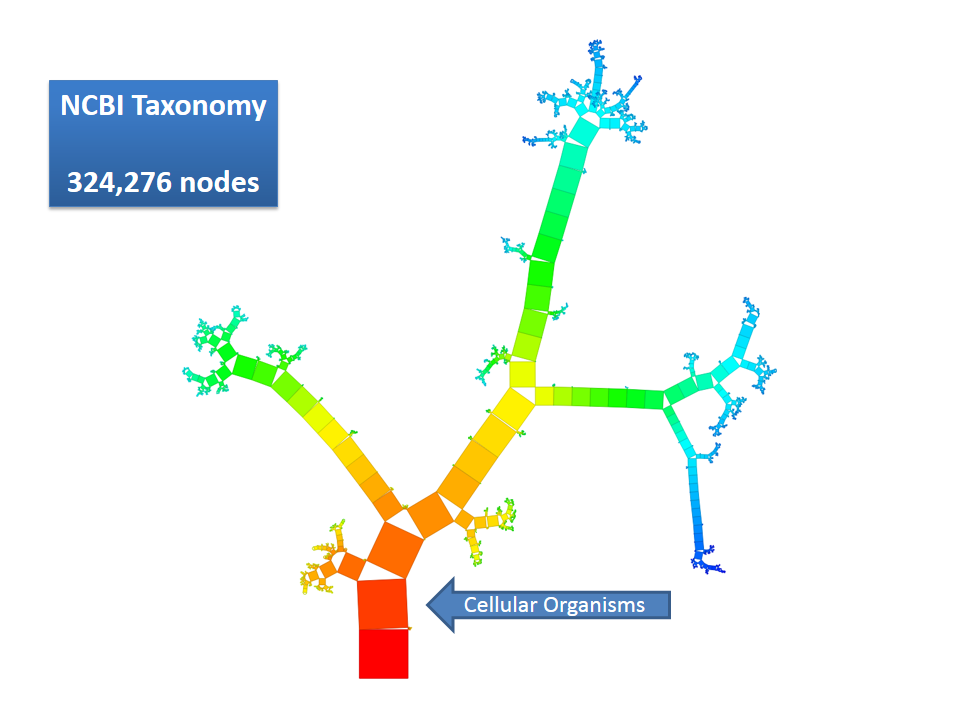
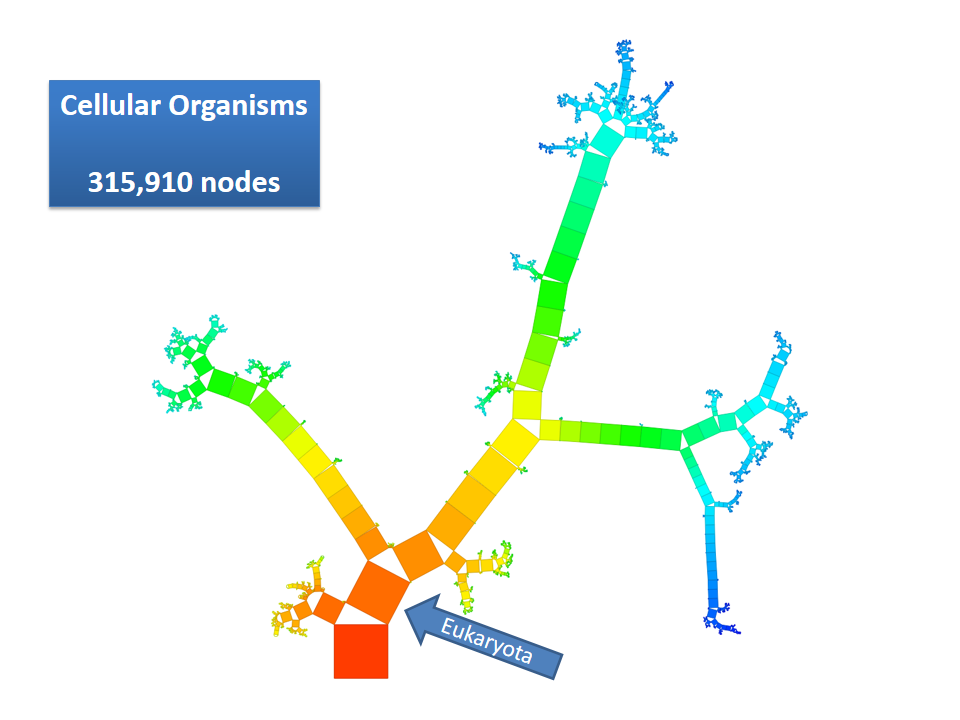
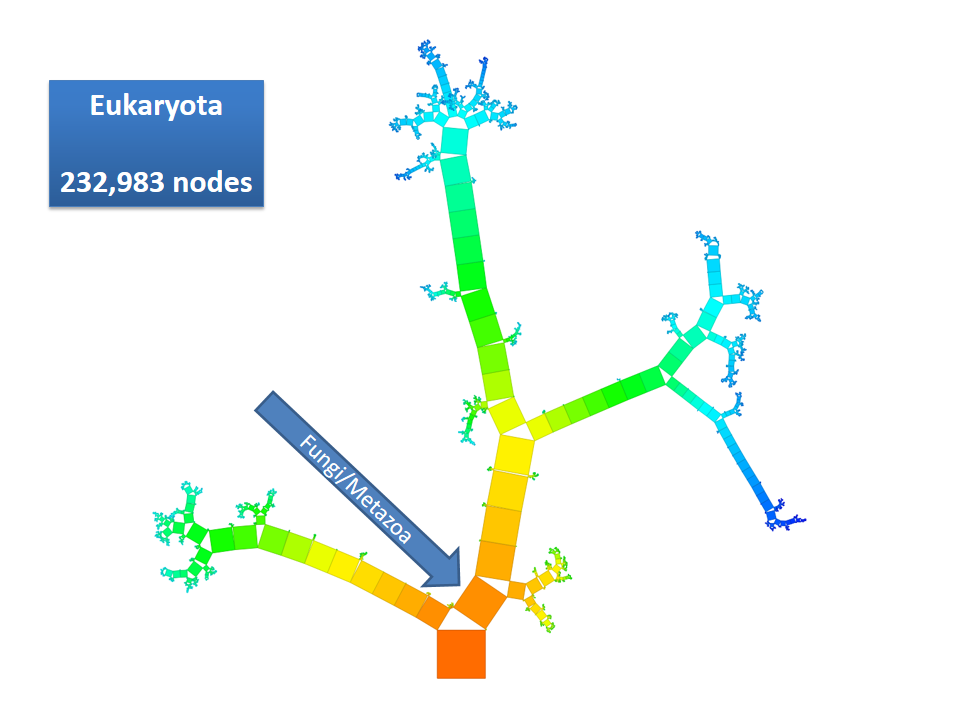
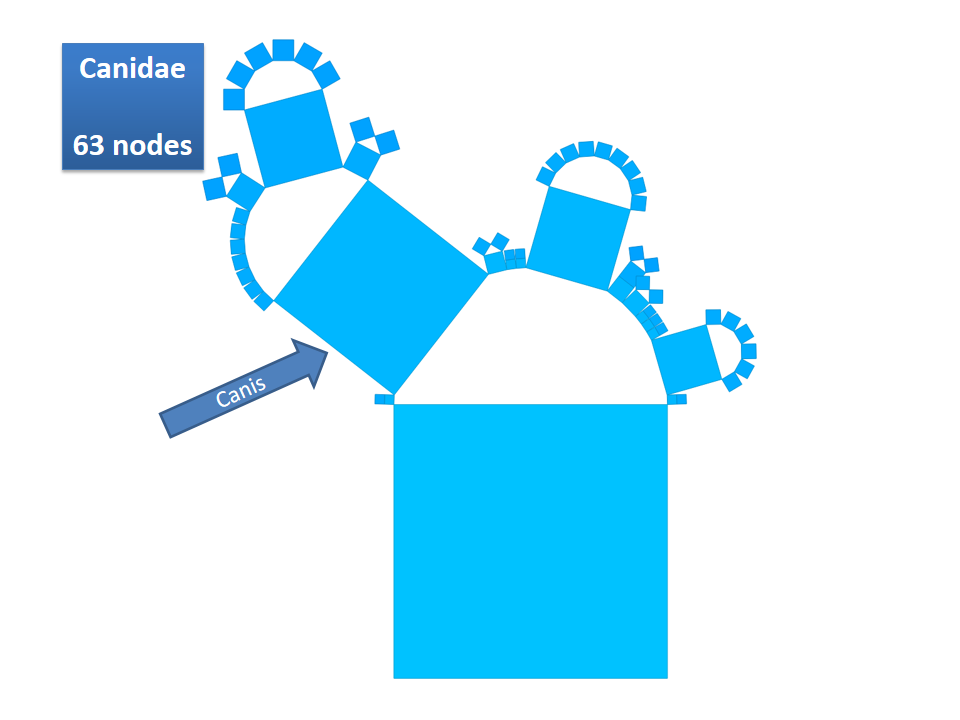
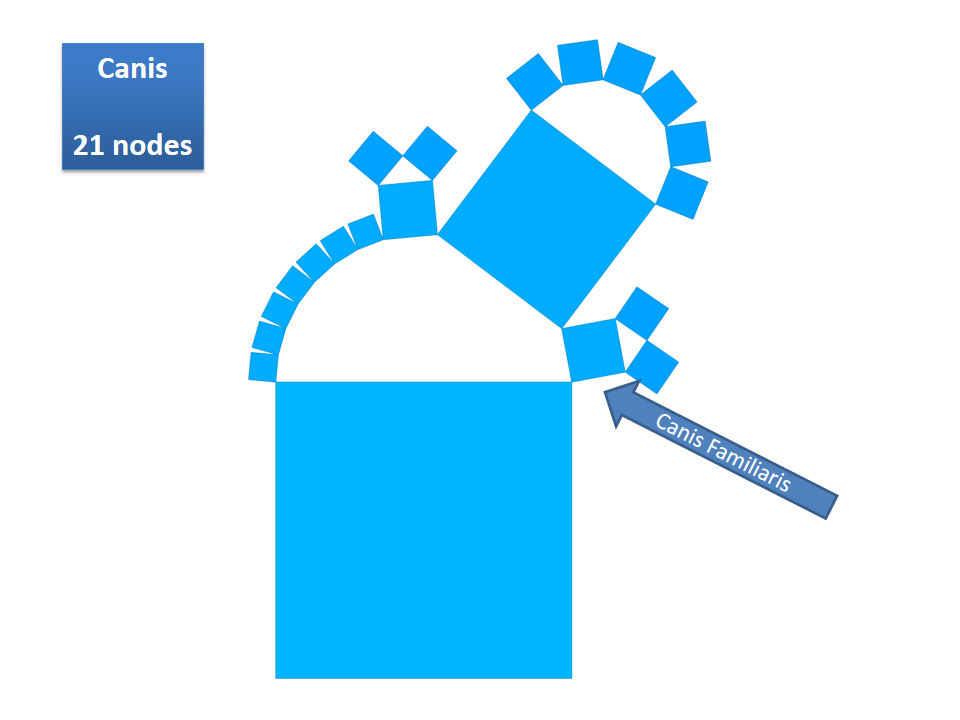
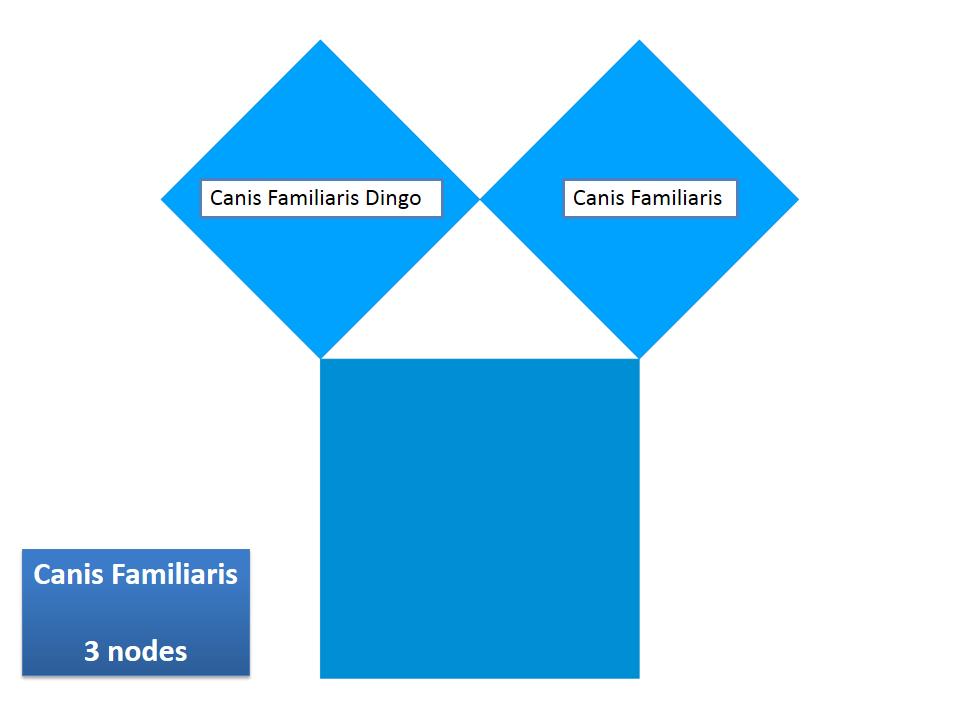
This example shows Generalized Pythagoras Trees applied to the NCBI tree of life. It contains 263,691 organisms classified into 60,585 groups of species (in total: 324,276 nodes). By clicking through the gallery you browse the tree down to a well-known animal: the common dog, Canis Familiaris.